IILLS
Viruses are the most abundant biological entity on the planet and widely distributed in the environments and organs of the world, and also an important part of the human microbiome which closely related with human health and disease. Viral infectious diseases are the important threat for human health, and receptor-binding is the first step for viral infection of host. In order to more effectively treat human viral diseases, it is needed to discover hidden virus-receptor interactions. Here, we developed a new virus-receptor interactions predictor IILLS , by using the Gaussian Interaction Profile (GIP) similarity to compute the similarity of viruses and the amino acid sequences similarity as the final similarity of receptors based on the experiment results. The IILLS provides two ways to input your data: a receptor sequence or a txt file with multiple sequences with FASTA format.
Read Me
Introduction
Viruses are the most abundant biological entity on the planet and widely distributed in the environments and organs of the world, and also an important part of the human microbiome which closely related with human health and disease. Viral infectious diseases are the important threat for human health, and receptor-binding is the first step for viral infection of host. In order to more effectively treat human viral diseases, it is needed to discover hidden virus-receptor interactions. The IILLS is a free web server for virus-receptor interaction prediction
Method
IILLS took the input receptor sequence in fasta format, either a pasted sequence or a file. Then the sequence similarity between input receptors is calculated by using GIP similarity calculating method and normalized Smith-Waterman method. In addition, IILLS uses Gaussian Interaction Profile (GIP) similarity for virus similarity measurement. Then, IILLS sets the initialized interaction scores for viruses (receptors) which have no known interaction with receptors (viruses) by KNN method. IILLS also uses LapRLS model to discover hidden virus-receptor interactions.
1 computing similarity of receptors
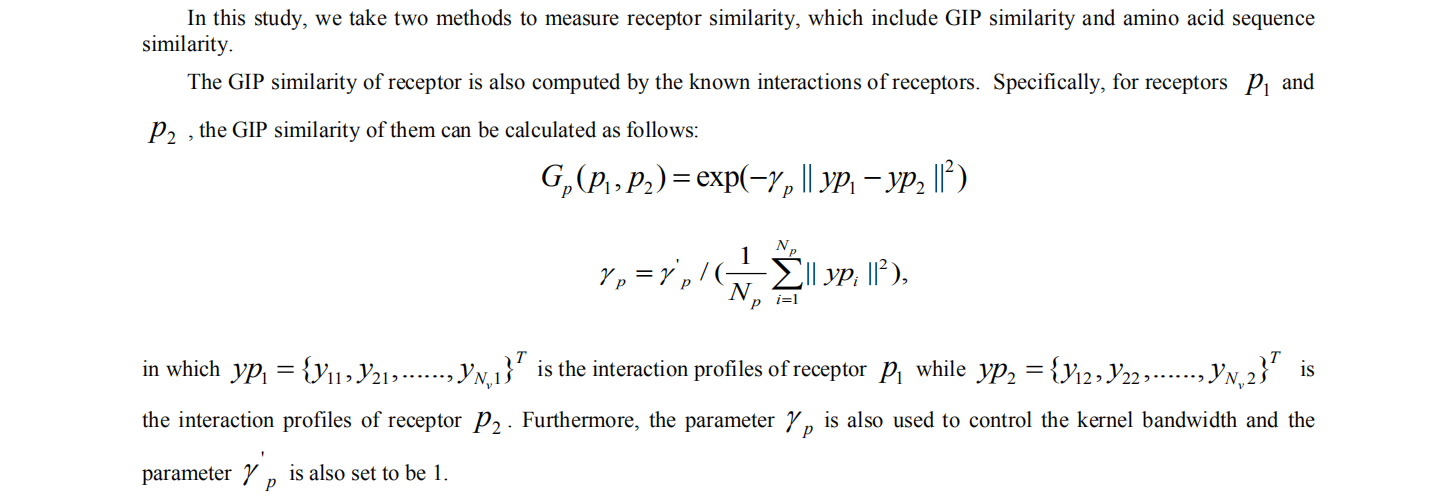
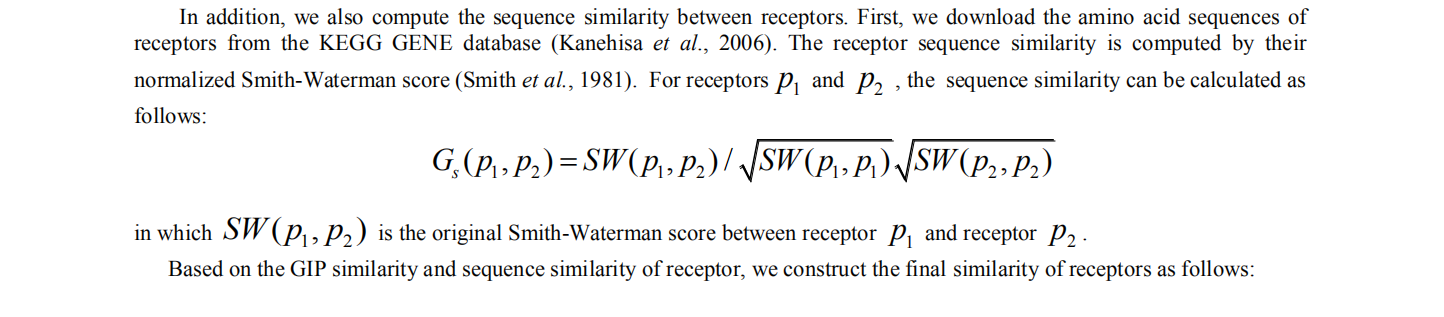
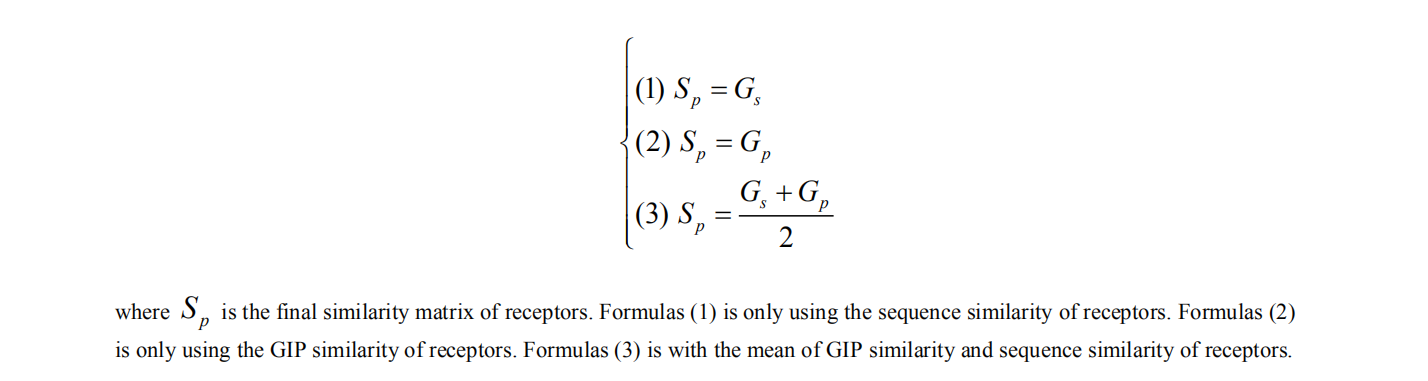



2. computing similarity of virus
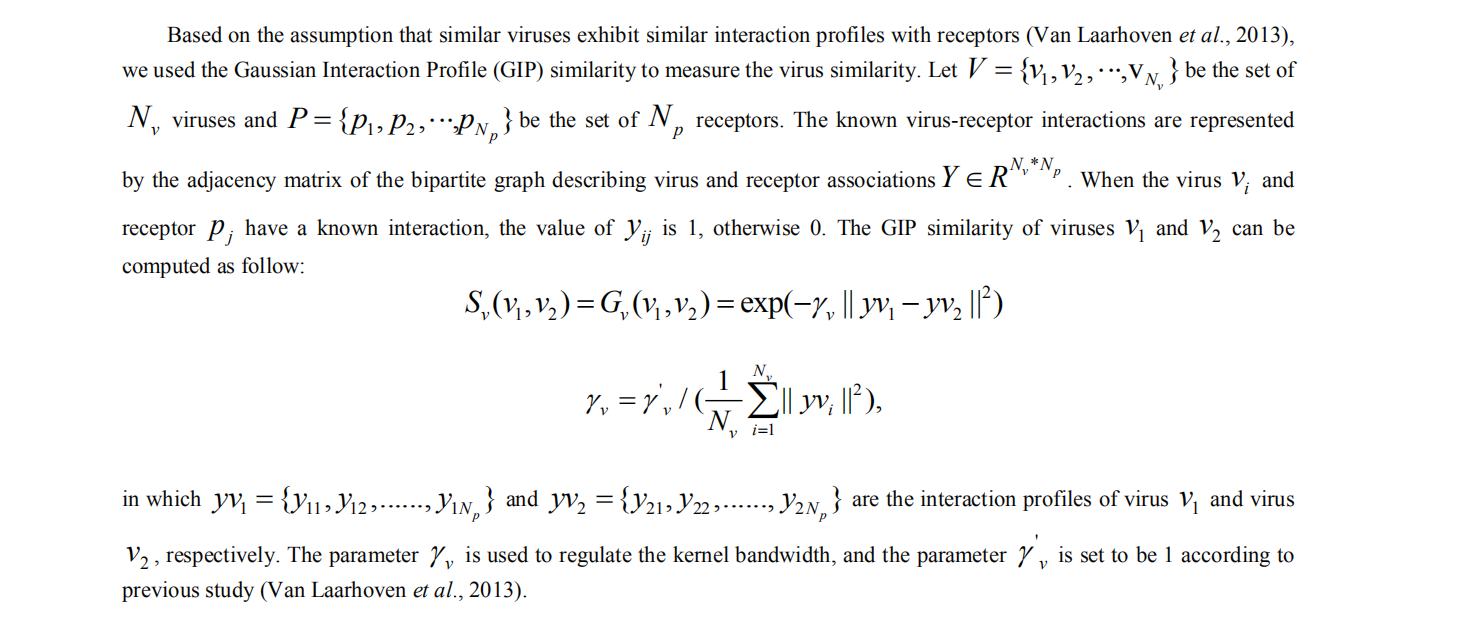

3. Initialized interaction scores for new viruses and receptors
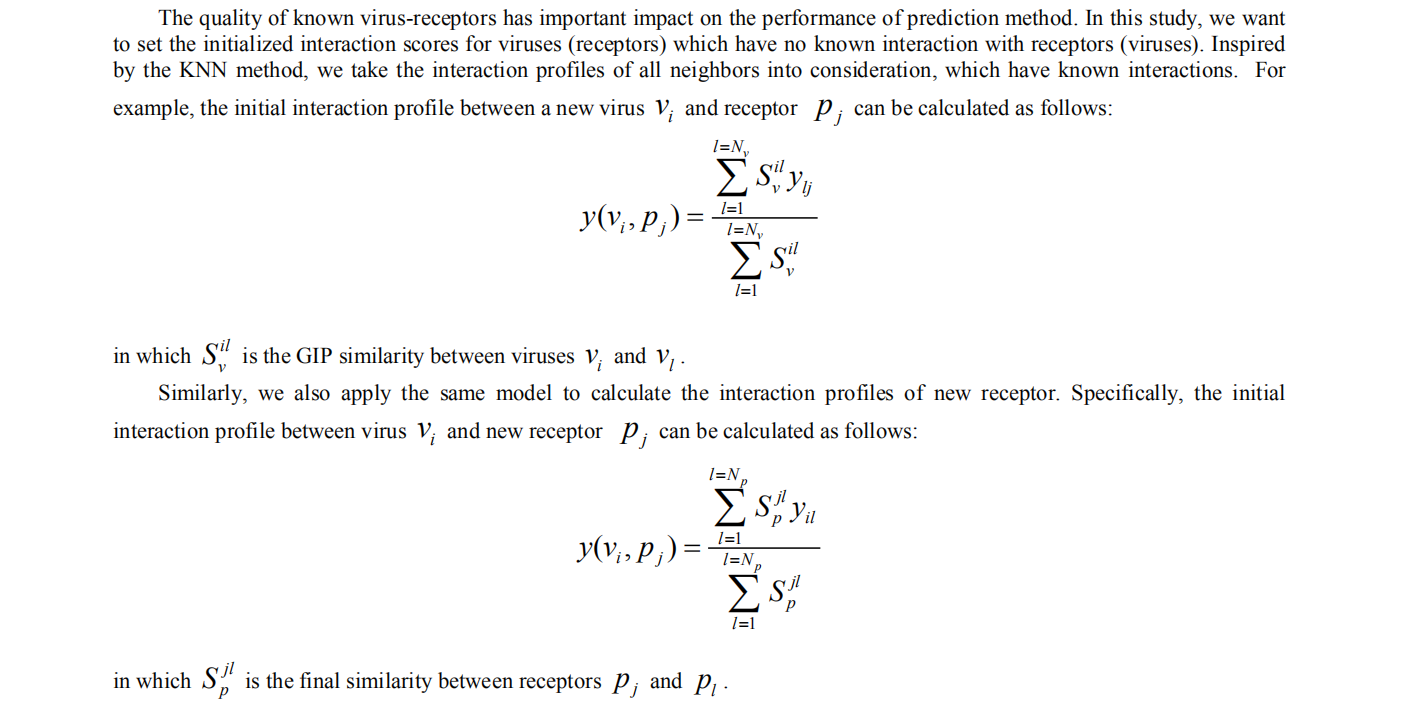
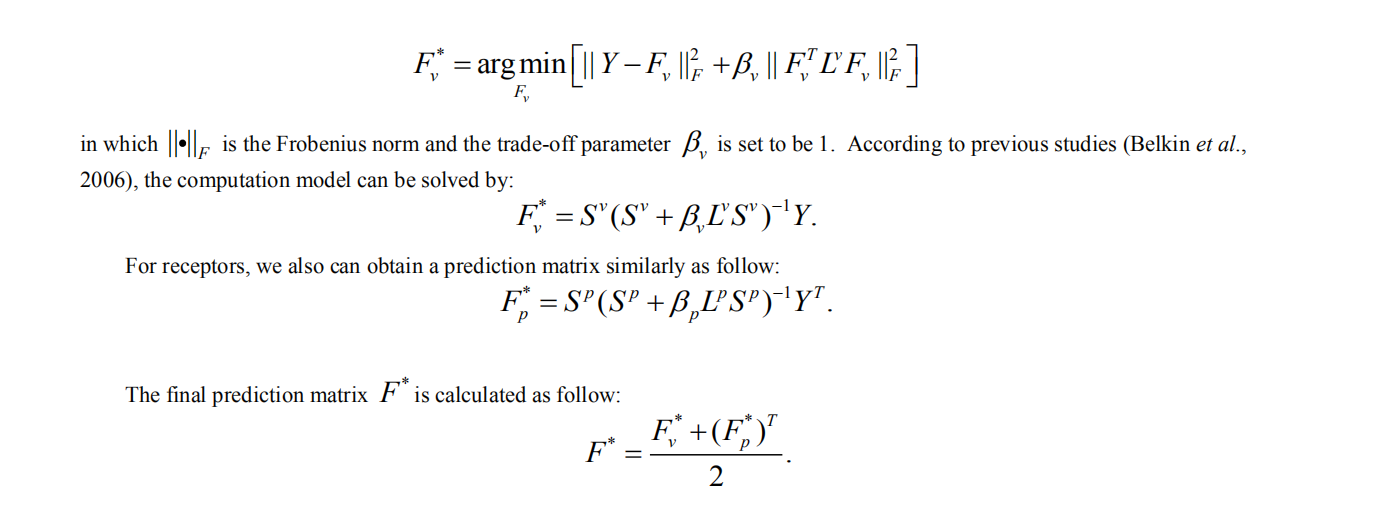


4. Laplacian regularized Least Square for virus-receptor interaction prediction
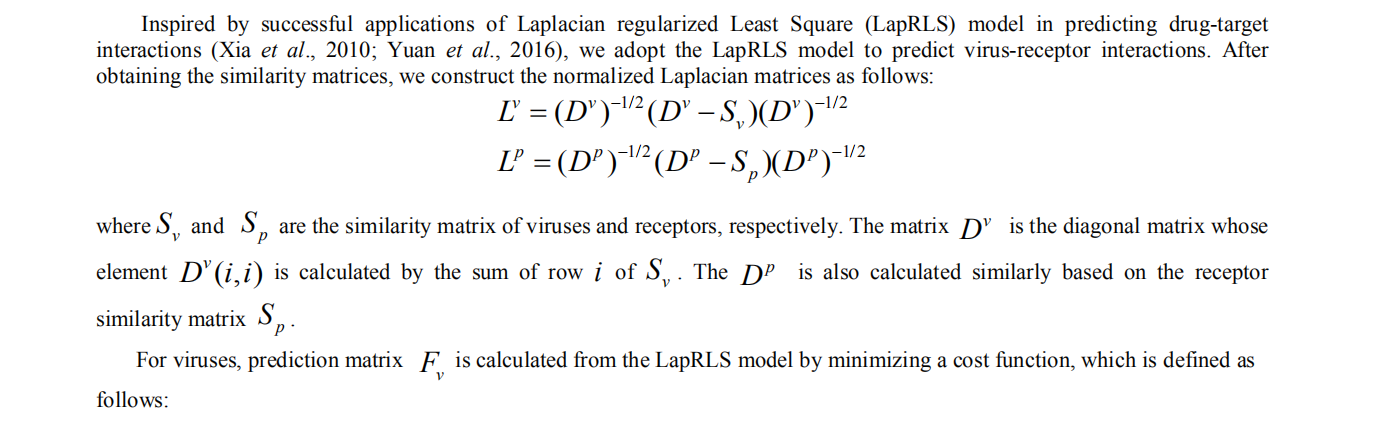
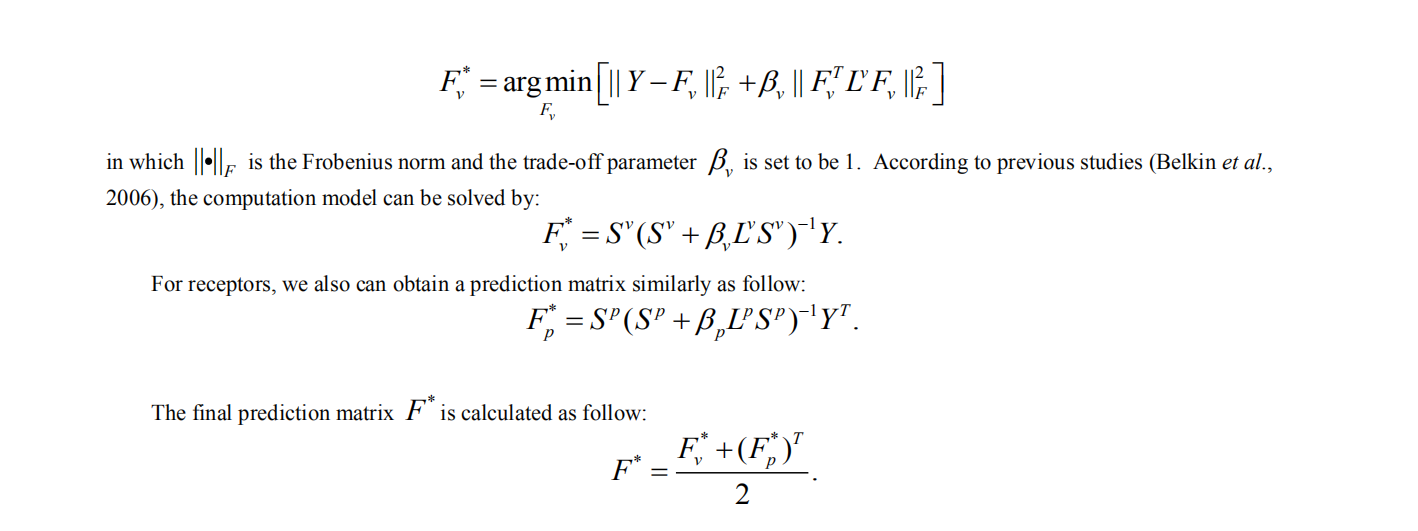


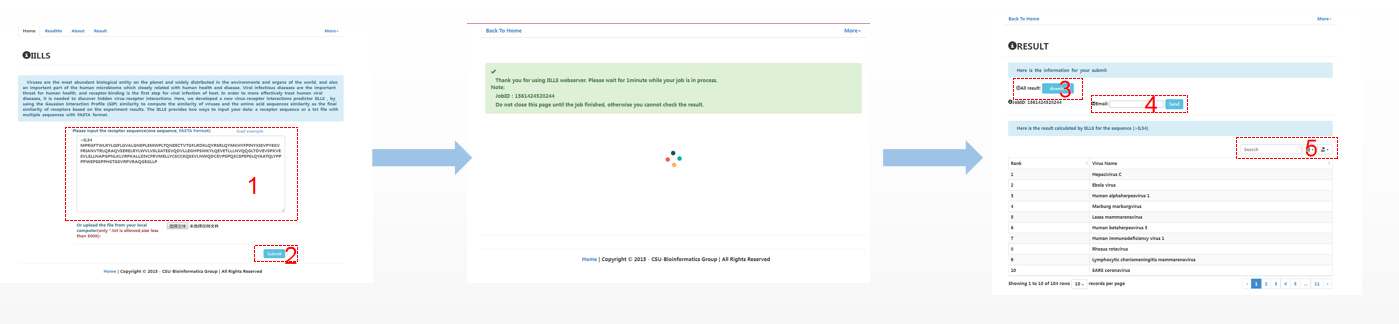
The IILLS took the input receptor sequence in fasta format, either a pasted sequence or a file with multiple sequences (size limit < 50kb). When the user want to submit single sequence, the user can paste the sequence with FASTA format into textbox (1) and click the “submit” button (2). The user could not close the page and waits for the result. The predicted result will be showed within several minutes. The user can download the results (3). If the user want to check the result in future, it can be regained by email (4). In addition, the user can search the result with virus name (5).
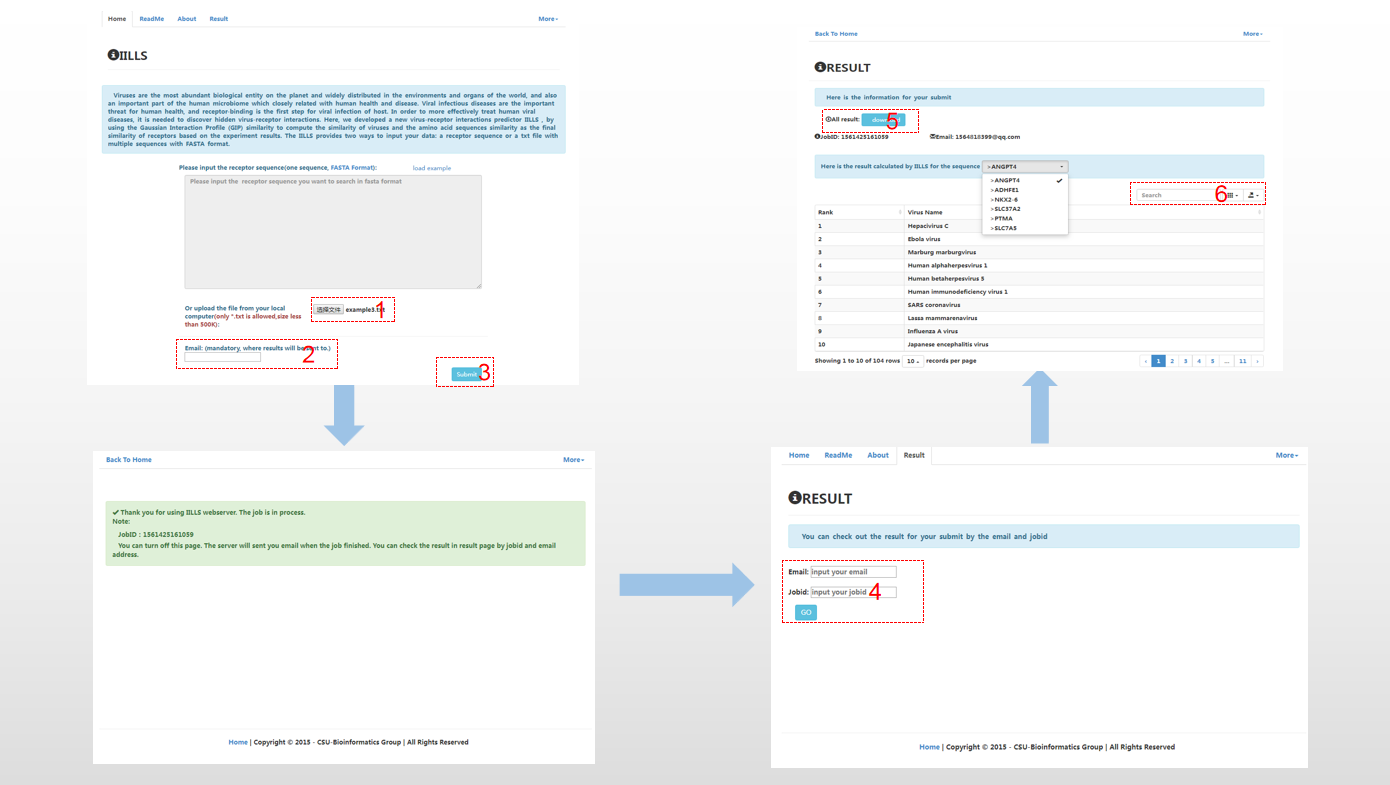
When the user submits text file with multiple sequences, the user uploads the text with multiple sequences (1) and provides email address (2). And then clicks the “submit” button (3), the user can close the page when submitting successfully. The notification will be sent to the user in term of email when task is completed. The user can check their result in result page by jobid and email (4). In addition, the user can download the results (5) and search the result with virus name (6).
Input Examples
The IILLS provides two ways to upload data:
1): single sequence
Single sequence: the user can paste their sequence with FASTA format (A sequence in FASTA format begins with a single-line description, followed by lines of sequence data. The description line is distinguished from the sequence data by a greater-than (">") symbol in the first column.). Three input examples of single sequence are listed below:
>IL34
MPRGFTWLRYLGIFLGVALGNEPLEMWPLTQNEECTVTGFLRDKLQYRSRLQYMKHYFPINYKISVPYEGVFRIANVTRLQRAQVSERELRYLWVLVSLSATESVQDVLLEGHPSWKYLQEVETLLLNVQQGLTDVEVSPKVESVLSLLNAPGPNLKLVRPKALLDNCFRVMELLYCSCCKQSSVLNWQDCEVPSPQSCSPEPSLQYAATQLYPPPPWSPSSPPHSTGSVRPVRAQGEGLLP
>OR5K1
MAEENHTMKNEFILTGFTDHPELKTLLFVVFFAIYLITVVGNISLVALIFTHRRLHTPMYIFLGNLALVDSCCACAITPKMLENFFSENKRISLYECAVQFYFLCTVETADCFLLAAMAYDRYVAICNPLQYHIMMSKKLCIQMTTGAFIAGNLHSMIHVGLVFRLVFCGSNHINHFYCDILPLYRLSCVDPYINELVLFIFSGSVQVFTIGSVLISYLYILLTIFKMKSKEGRAKAFSTCASHFLSVSLFYGSLFFMYVRPNLLEEGDKDIPAAILFTIVVPLLNPFIYSLRNREVISVLRKILMKK
>GAPDHS
MSKRDIVLTNVTVVQLLRQPCPVTRAPPPPEPKAEVEPQPQPEPTPVREEIKPPPPPLPPHPATPPPKMVSVARELTVGINGFGRIGRLVLRACMEKGVKVVAVNDPFIDPEYMVYMFKYDSTHGRYKGSVEFRNGQLVVDNHEISVYQCKEPKQIPWRAVGSPYVVESTGVYLSIQAASDHISAGAQRVVISAPSPDAPMFVMGVNENDYNPGSMNIVSNASCTTNCLAPLAKVIHERFGIVEGLMTTVHSYTATQKTVDGPSRKAWRDGRGAHQNIIPASTGAAKAVTKVIPELKGKLTGMAFRVPTPDVSVVDLTCRLAQPAPYSAIKEAVKAAAKGPMAGILAYTEDEVVSTDFLGDTHSSIFDAKAGIALNDNFVKLISWYDNEYGYSHRVVDLLRYMFSRDK
2): Text file with multiple sequences
Text file with multiple sequences: if the user want to upload
multiple sequences, we recommend the user upload their data with
text file. In text file, each sequence should be the FASTA format.
The size of text file should be 50KB. The input example of text is
listed below (In package, it can four text files: example1.txt,
example2.txt, example3.txt and example4.txt contains three, five, six, four
receptor sequences, respectively.):
Output Description
The final results will be given in a table format. Each sequence id follows the result of predicted virus with scores ranging from 0 to 1. Higher scores represent higher confidence of the virus. Downloadable results are provided in txt format. If email address is left previously, an alert mail will be sent as soon as the results are ready.
Table 1 shows the validation results of top 10 virus-receptor interactions of the entire dataset which are predicted by IILLS. C-type lectin domain family 4 member M (CLEC4M, also called L-SIGN or CD209L) is equipped with a carbohydrate recognition domain (CRD) that mediates the recognition of fucose and high-mannose glycans in a Ca2+-dependent manner, these carbohydrate structures can be found in multiple pathogens, such as Lassa virus, Ebola virus, among others (Garcia-Vallejo et al., 2015 and Sakuntabhai et al., 2005). The CD209 molecule (CD209) is the receptors of diseases which caused by the previously known human coronaviruses, Human coronavirus 229E (229E) (Lo et al., 2006). L-SIGN also called DC-SIGN related (CLEC4M) is C-type lectin involved in both innate and adaptive immunity, they are the known to bind multiple pathogens and function as cellular receptors for various viruses, such as Dengue virus (Li et al., 2012). Rift Valley fever virus (RVFV) used L-SIGN to infect cells expressing the lectin ectopically (Léger et al., 2016, and Sakuntabhai et al., 2005). The phleboviruses, such as Uukuniemi virus (UUKV), can exploit L-SIGN for infection (Léger et al., 2016, and Sakuntabhai et al., 2005).
| Virus | Receptor | References | |
|---|---|---|---|
| Lymphocytic choriomeningitis mammarenavirus (LCMV) | C-type lectin domain family 4 member M(CLEC4M, L-SIGN) | Unknown | |
| Lassa mammarenavirus | C-type lectin domain family 4 member M | Garcia-Vallejo et al, (2015) and Sakuntabhai et al., (2005) | |
| Human coronavirus 229E(229E) | CD209 molecule (CD209) | Lo et al., (2006) | |
| Dengue virus | C-type lectin domain family 4 member M | Li et al., (2012) | |
| Rift Valley fever virus | C-type lectin domain family 4 member M | Léger et al., (2016), and Sakuntabhai et al., (2005) | |
| Uukuniemi virus | C-type lectin domain family 4 member M | Léger et al., (2016), and Sakuntabhai et al., (2005) | |
| Human immunodeficiency virus 2 | C-type lectin domain family 4 member M | Unknown | |
| Human alphaherpesvirus 1 | integrin subunit beta 3 (beta 3 integrin) | Unknown | |
| Coxsackievirus A9(CAV9) | integrin subunit beta 1 | Unknown | |
| Human betaherpesvirus 5 | integrin subunit beta 6 | Unknown |
About
IILLS is a non-profit tool to provide web severs for predictimg virus-receptor interactions.
Questions and comments on the tool and suggestions for improvement to the website are always welcome.
Please contact:
Cheng Yan: yancheng01@mail.csu.edu.cn
Guihua Duan:duangh@mail.csu.edu.cn
Jianxin Wang: jxwang@mail.csu.edu.cn
Fang-xiang Wu: faw341@mail.usask.ca
Result
You can check out the result for your submit by the email and jobid